Difference between revisions of "BLAST"
From Pheno Wiki
| Line 1: | Line 1: | ||
== BLAST - Basic Logical Alignment Search Tool == | == BLAST - Basic Logical Alignment Search Tool == | ||
| − | '''What | + | '''What it does:''' |
| Line 13: | Line 13: | ||
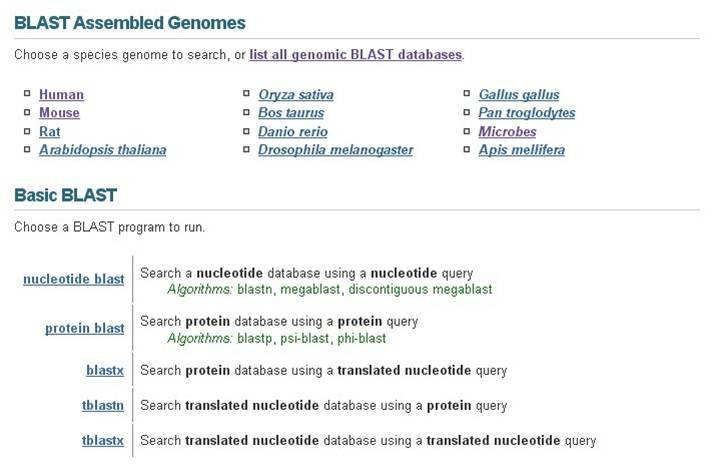
'''Main Page''' | '''Main Page''' | ||
[[File:Blast_main_page.jpg]] | [[File:Blast_main_page.jpg]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | '''Example Use:''' | ||
| + | You’ve recently cloned a mouse adrenergic receptor, but you want to know if there are any homologous sequences in the mouse genome that might code for other receptor isoforms | ||
| + | |||
| + | Why not BLAST sequence from your cloned receptor… | ||
| + | |||
| + | |||
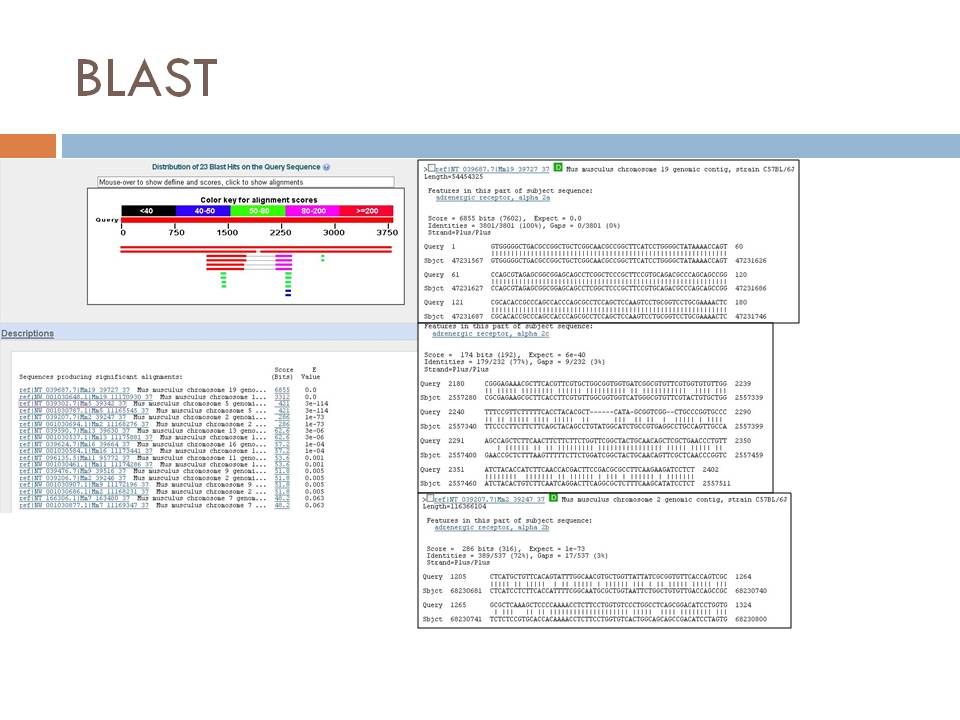
| + | '''BLAST results''' | ||
| + | [[File:Blast_results.jpg]] | ||
Revision as of 14:31, 18 January 2010
BLAST - Basic Logical Alignment Search Tool
What it does:
“…finds regions of local similarity between sequences.”
“…compares nucleotide or protein sequences to sequence databases and calculates the statistical significance of matches.”
“… can be used to infer functional and evolutionary relationships between sequences as well as help identify members of gene families.”
Example Use:
You’ve recently cloned a mouse adrenergic receptor, but you want to know if there are any homologous sequences in the mouse genome that might code for other receptor isoforms
Why not BLAST sequence from your cloned receptor…