Quality Control
Back to LA5C
Contents
Check setup_subject
To make sure everything ran correctly check:
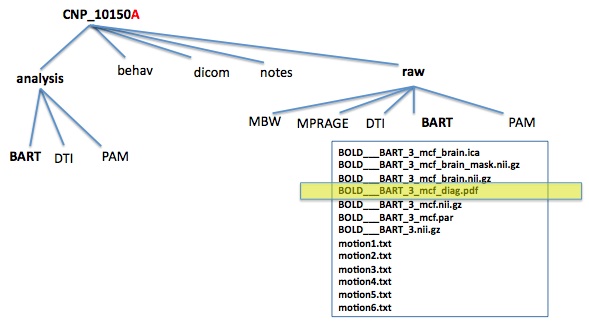
- directory structure
- files
- upload behavior files
For details on setup subjects go here: Initial_Processing
Inspect Diagnostic Reports
- Flag excessive motion
- Diagnostic Reports
- Stored in the raw/TASK folder for each task
- Use display or kpdf to open it
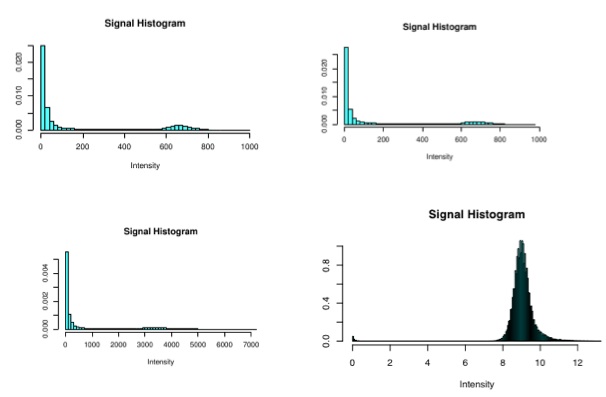
Diagnostic Examples: Histograms
These histograms show the range of signal in the raw image. It is normal to have a large peak centered around 0 that represents the background, or non-brain voxels. If this is missing, or low, then there may be an issue with background noise. It is then normal to have another, much lower, peak farther out centered around 800-1200, that represents the data in the brain.
In these examples, you can see that the obviously strange looking one on the lower right hand side is from a scan that was entirely noise- this is a very abnormal example.
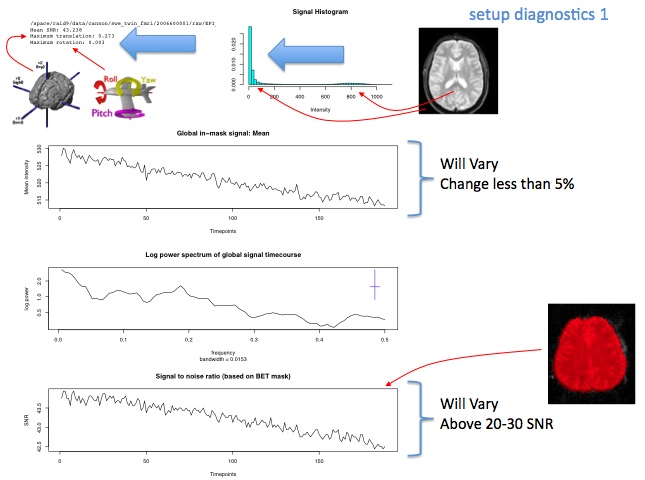
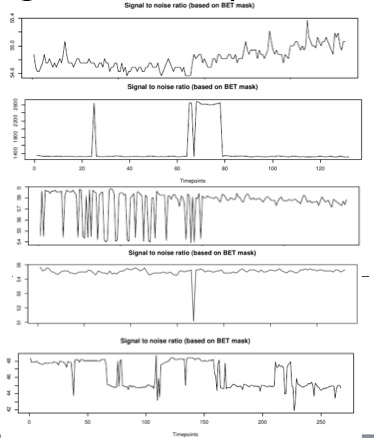
Diagnostic Examples: SNR
This measure is calculated based on the binarized mask created when BET is run.
For SNR, there are a few things to check. First of all you want to be sure it is at least above 20-30. Lower than that indicates very low signal. Second, it is important to check for large fluctuations (for instance, if there is substantial instability at the start or end of the scan). Often, something that looks like a large fluctuation is actually not relevant when you look at the y-axis and see that it actually represents a very small percent change. This axis re-scales itself based on the data, so you must check it in order to interpret the findings. For instance, example 2 below looks similar to the others, but the scale is very large, and this is likely significant, where in other similar looking graphs with smaller ranges, it would not be.
Sometimes there will be a spike in the exact center of the graph. This likely represents an artifact based on fat suppression. The motion correction is based on the center image, so all images other than the center image have been interpolated to some extent. If fat wasn't sufficiently suppressed, the middle image may have much lower SNR because in all the other images the higher intensity fat has been blurred around as the correction occurred. The decrease in SNR seems to be totally based on an increase in background noise, not on any decrease within the brain and the within brain data will be fine. This is not usually relevant, and will be common across subjects within a protocol (see example 3, below). You can detect it by checking other subjects on the same protocol, and by checking to see if it is happening precisely on the middle scan.
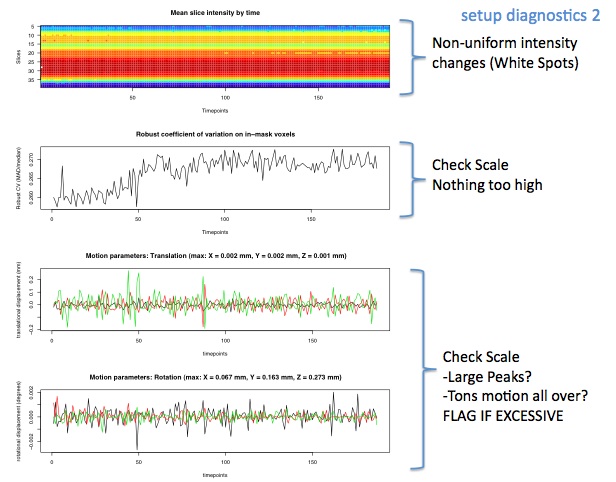
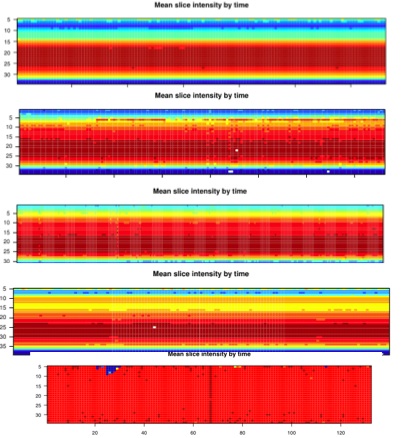
Diagnostic Examples: Slice Intensity
This image represents the signal intensity across each time point on each slice. There is normally a thick red band that represents the area with the most data, which for us is the head. There are sometimes white dots in this- if there are only a few, it should be noted, but doesn't usually correlate with significant problems with the data. If the pattern is messy, or blurry, or the red band is not particularly wide or dark, then this can often correlate with motion, or otherwise bad data. This image in and of itself is not sufficient evidence for eliminating subjects, but can help you understand the impact other parts of the diagnostics (like motion, or SNR) are having on your data.
The top image is an example of a normal scan.
The middle images are in descending order of messiness.
The bottom image is entirely noise (it is the same as the example of the bad histogram, above). You can see that there is no specificity in terms of head location, since the entire thing is red.
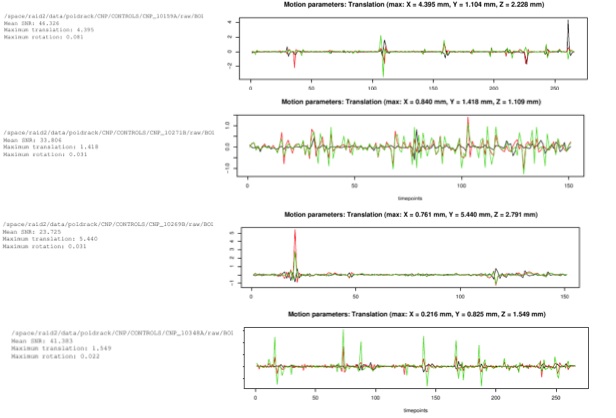
Diagnostic Examples: Motion
Motion is probably the most important component of data checking, as it can really ruin the data, especially with interleaved scans. Usually, anything over 2-3 mm translation is noteworthy, and potentially warrants exclusions.
When checking motion, be sure to look at the y-axis. This graph also re-scales itself, so it can be misleading if you don't pay attention to scale. It is often useful to look at how the motion spikes here line up with artifacts seen in the previous graphs as it will often line up with disruptions in things like the color map.
Any unusual motion should be logged. Generally, spikes in motion are much more disruptive than a slow drift, which can be more easily modelled out by FSL.
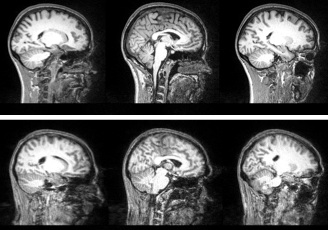
Diagnostic Examples: MPRAGE
Good
Bad
Diagnostic Criteria
| Parameter | Mean range | Features |
|---|---|---|
| Global in-mask signal: Mean | 4-12 | gradual decline/overall negative slope |
| Log power spectrum of global signal timecourse | 2-9 | gradual decline/negative slope, long wave |
| Signal to noise (SNR) ration (based on BET mask) | 2-8 | no isolated spikes |
| Mean slice intensity by time | range of colors, no white spots, no white lines, no large bands of color | |
| Motion parameters: Translation | 0.2-1.2mm | no large spikes that extend the overall range by >1mm, flag any spike >2mm or >3mm |
Inspect raw data with fslview
- Number of scans
- In fslview
- fslvols <.nii file>
- Motion
- Orientation
- Brain extraction
- The arrow at the top emphasizes the film strip, clicking plays a video of slices (check for motion).
- The arrow at the bottom emphasizes volume, allowing you to select specific slices.
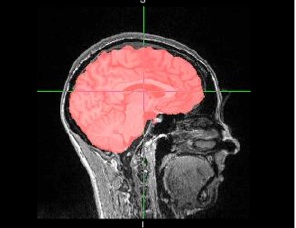
Inspect BET
- Usage: bet <input fileroot> <output fileroot> [options] example: bet spt100hr.hdr spt100hr_brain.hdr
- How it works: Searches out from center, looking for area of decreased signal, and cuts off there (knowing this will help a lot with understanding where artifacts come from).
- Run, then check using slices. If needs fixing:
- -f, fractional intensity threshold, ranges from 0-1. The default is .5, and making this number smaller causes the resulting brain to be bigger, while making it larger (>.5) makes the resulting brain smaller.
- -g, vertical gradient in fractional intensity threshold, ranges from -1 - 1. The default is 0. Closer to 1 makes brain have smaller top, bigger bottom. Closer to -1 makes brain have smaller bottom, bigger top (so select based on which part of brain is not being correctly estimated).
- example: bet spt100hr.hdr spt100hr.brain.hdr -f 0.4 –g .2
BET examples
- MPRAGE after setup_subject default bet
- Usage : > bet mprage.nii.gz mprage_brain –B –f 0.20
Details on Skull Stripping with BET can be found here: Skull_Strip_with_BET
Link back to LA5C page.