Difference between revisions of "CNP VMNM"
(→Code/Algorithms) |
(→Cleaning Rules) |
||
| Line 16: | Line 16: | ||
===Cleaning Rules=== | ===Cleaning Rules=== | ||
| − | 1. If any of the calculated variables are missing, that subject should be listed for exclusion. | + | 1. If any of the calculated variables are missing, that subject should be listed for exclusion.<br/> |
| − | 2. If Trial Count is not =40, then that subject should be listed for exclusion. | + | 2. If Trial Count is not =40, then that subject should be listed for exclusion.<br/> |
| − | 3. If overall performance (average percent correct) is below 50% (chance, in this task), that subject should be listed for exclusion. | + | 3. If overall performance (average percent correct) is below 50% (chance, in this task), that subject should be listed for exclusion.<br/> |
| − | 4. If average RT is greater than 6000ms that subject should be listed for exclusion. | + | 4. If average RT is greater than 6000ms that subject should be listed for exclusion.<br/> |
| − | It is worth considering dropping people with performance >50% on either maintenance or manipulation (rather than just on the average). | + | It is worth considering dropping people with performance >50% on either maintenance or manipulation (rather than just on the average).<br/> |
If you want to know if your group looks approximately like it should, in every sample run so far, accuracy is lower in manipulation than maintenance, and reaction time is greater in manipulation than maintenance. This should be the first group level analysis, to quickly verify everything is correct. It would be highly unexpected to have a deviation from this pattern, even though the position along the y axis (overall accuracy or RT) may vary, especially in patient groups. | If you want to know if your group looks approximately like it should, in every sample run so far, accuracy is lower in manipulation than maintenance, and reaction time is greater in manipulation than maintenance. This should be the first group level analysis, to quickly verify everything is correct. It would be highly unexpected to have a deviation from this pattern, even though the position along the y axis (overall accuracy or RT) may vary, especially in patient groups. | ||
Revision as of 17:45, 27 June 2011
go back to HTAC
Contents
Basic Task Description
Task Procedure
Task Structure Detail
Task Schematic
Task Parameters Table
Stimuli
Dependent Variables
Cleaning Rules
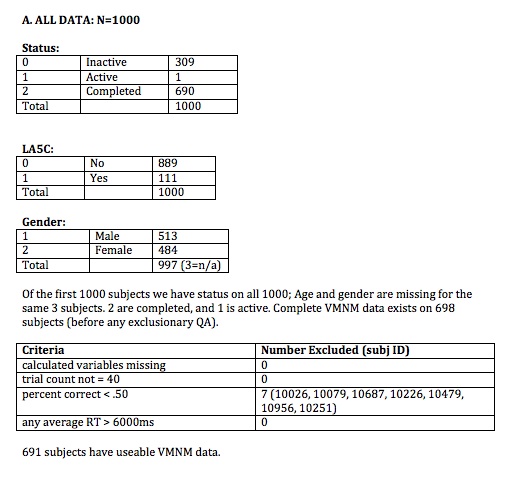
1. If any of the calculated variables are missing, that subject should be listed for exclusion.
2. If Trial Count is not =40, then that subject should be listed for exclusion.
3. If overall performance (average percent correct) is below 50% (chance, in this task), that subject should be listed for exclusion.
4. If average RT is greater than 6000ms that subject should be listed for exclusion.
It is worth considering dropping people with performance >50% on either maintenance or manipulation (rather than just on the average).
If you want to know if your group looks approximately like it should, in every sample run so far, accuracy is lower in manipulation than maintenance, and reaction time is greater in manipulation than maintenance. This should be the first group level analysis, to quickly verify everything is correct. It would be highly unexpected to have a deviation from this pattern, even though the position along the y axis (overall accuracy or RT) may vary, especially in patient groups.
Code/Algorithms
Calculation rules are all the same as above.
This is the code for data QA, in Stata.
//ALL SUBJECTS
//get breakdown of numbers in each group:
bysort status: summarize ptid
bysort la5ccontrol: summarize ptid
bysort gender: summarize ptid
//see how many complete cases there are.
count if vmnm_main_mn !=.
//exclusions
generate float vmnm_percent_corr=(vmnm_manip_mn + vmnm_main_mn)/2
count if vmnm_percent_corr<=.5
count if vmnm_trialcount != 40 & vmnm_main_tt !=.
count if (vmnm_main_mnrt >6000 | vmnm_manip_mnrt > 6000) & vmnm_main_tt !=.
//create variable summing exclusion criteria
generate float useable_vmnm= 1 if vmnm_trialcount==40 & vmnm_percent_corr>.5 & vmnm_main_mnrt< 6000 & vmnm_manip_mnrt<6000
count if useable_vmnm==1
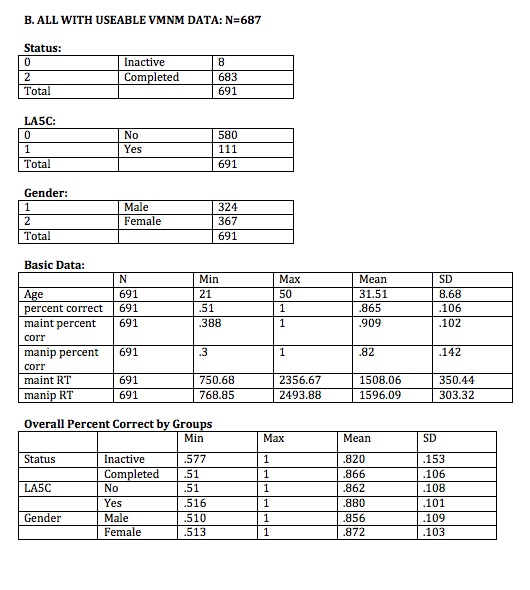
//SUBJECTS WITH USEABLE DATA
bysort status: summarize ptid if useable_vmnm==1
bysort la5ccontrol: summarize ptid if useable_vmnm==1
bysort gender: summarize ptid if useable_vmnm==1
summarize age vmnm_percent_corr vmnm_main_mn vmnm_manip_mn vmnm_main_mnrt vmnm_manip_mnrt if useable_vmnm==1
bysort status: summarize vmnm_percent_corr vmnm_main_mn vmnm_manip_mn vmnm_main_mnrt vmnm_manip_mnrt if useable_vmnm==1
bysort la5ccontrol: summarize vmnm_percent_corr vmnm_main_mn vmnm_manip_mn vmnm_main_mnrt vmnm_manip_mnrt if useable_vmnm==1
bysort gender: summarize vmnm_percent_corr vmnm_main_mn vmnm_manip_mn vmnm_main_mnrt vmnm_manip_mnrt if useable_vmnm==1
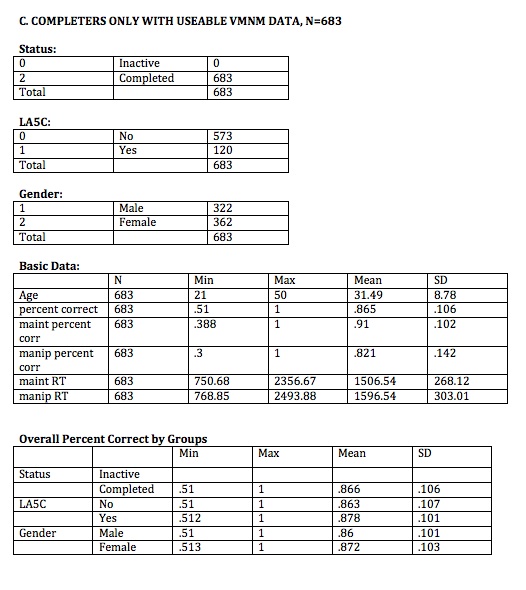
//COMPLETERS WITH USEABLE DATA
generate float comp_useable_vmnm= 1 if vmnm_trialcount==40 & vmnm_percent_corr>.5 & vmnm_main_mnrt< 6000 & vmnm_manip_mnrt<6000 &status==2
count if comp_useable_vmnm==1
bysort status: summarize ptid if comp_useable_vmnm==1
bysort la5ccontrol: summarize ptid if comp_useable_vmnm==1
bysort gender: summarize ptid if comp_useable_vmnm==1
summarize age vmnm_percent_corr vmnm_main_mn vmnm_manip_mn vmnm_main_mnrt vmnm_manip_mnrt if comp_useable_vmnm==1
bysort status: summarize vmnm_percent_corr vmnm_main_mn vmnm_manip_mn vmnm_main_mnrt vmnm_manip_mnrt if comp_useable_vmnm==1
bysort la5ccontrol: summarize vmnm_percent_corr vmnm_main_mn vmnm_manip_mn vmnm_main_mnrt vmnm_manip_mnrt if comp_useable_vmnm==1
bysort gender: summarize vmnm_percent_corr vmnm_main_mn vmnm_manip_mn vmnm_main_mnrt vmnm_manip_mnrt if comp_useable_vmnm==1
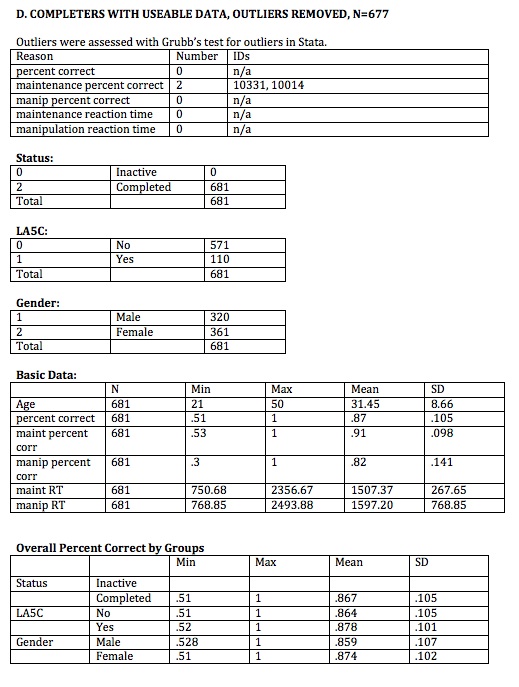
//OUTLIERS
sktest vmnm_percent_corr vmnm_main_mn vmnm_manip_mn vmnm_main_mnrt vmnm_manip_mnrt
grubbs vmnm_manip_mn if comp_useable_vmnm==1, gen (grubb_vmnm_manip_mn)
grubbs vmnm_main_mn if comp_useable_vmnm==1, gen(grubb_vmnm_main_mn)
grubbs vmnm_manip_mnrt if comp_useable_vmnm==1, gen (grubb_vmnm_manip_mnrt)
grubbs vmnm_main_mnrt if comp_useable_vmnm==1, gen(grubb_vmnm_main_mnrt)
grubbs vmnm_percent_corr if comp_useable_vmnm==1, gen(grubb_vmnm_percorr)
generate float comp_useable_nooutlier_vmnm=1 if comp_useable_vmnm==1 & grubb_vmnm_manip_mn !=1 & grubb_vmnm_main_mn !=1 & grubb_vmnm_manip_mnrt !=1 & grubb_vmnm_main_mnrt !=1 & grubb_vmnm_percorr !=1
count if comp_useable_nooutlier_vmnm==1
bysort status: summarize ptid if comp_useable_nooutlier_vmnm==1
bysort la5ccontrol: summarize ptid if comp_useable_nooutlier_vmnm==1
bysort gender: summarize ptid if comp_useable_nooutlier_vmnm==1
summarize age vmnm_percent_corr vmnm_main_mn vmnm_manip_mn vmnm_main_mnrt vmnm_manip_mnrt if comp_useable_nooutlier_vmnm==1
bysort status: summarize vmnm_percent_corr vmnm_main_mn vmnm_manip_mn vmnm_main_mnrt vmnm_manip_mnrt if comp_useable_nooutlier_vmnm==1
bysort la5ccontrol: summarize vmnm_percent_corr vmnm_main_mn vmnm_manip_mn vmnm_main_mnrt vmnm_manip_mnrt if comp_useable_nooutlier_vmnm==1
bysort gender: summarize vmnm_percent_corr vmnm_main_mn vmnm_manip_mn vmnm_main_mnrt vmnm_manip_mnrt if comp_useable_nooutlier_vmnm==1
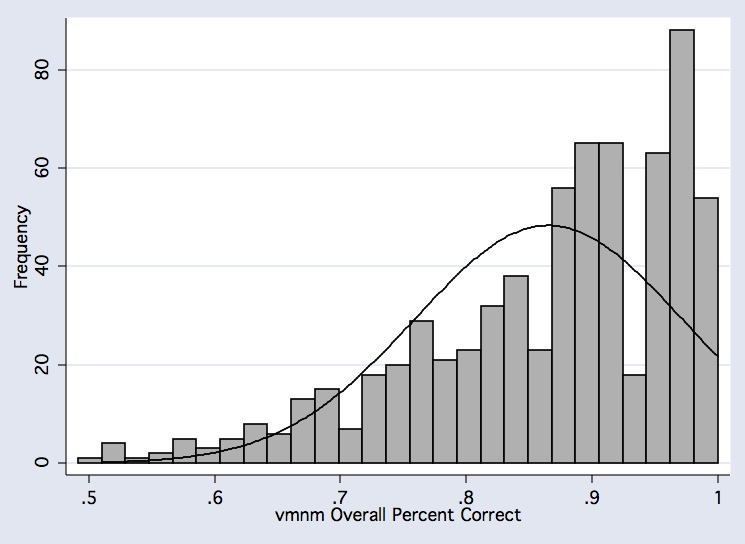
histogram vmnm_percent_corr if comp_useable_vmnm==1, frequency bfcolor(gs11) blcolor(black) normal normopts( clcolor(black) ) xtitle(vmnm Overall Percent Correct)
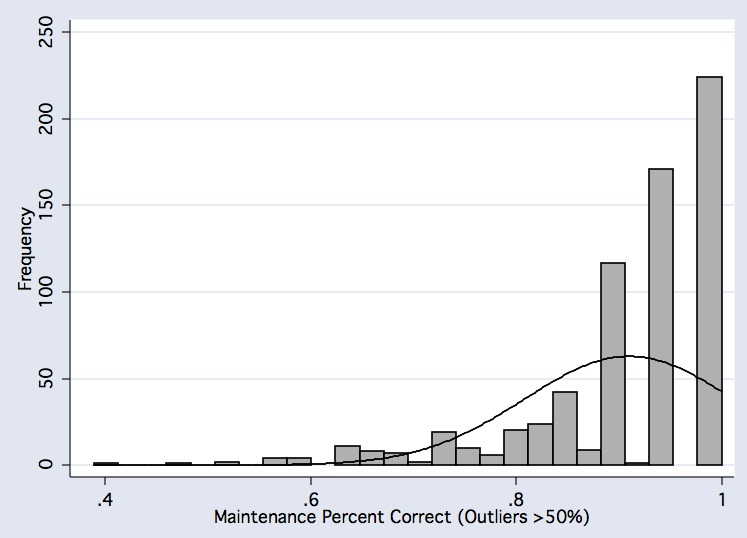
histogram vmnm_main_mn if comp_useable_vmnm==1, frequency bfcolor(gs11) blcolor(black) normal normopts( clcolor(black) ) xtitle(Maintenance Percent Correct (Outliers >50%))
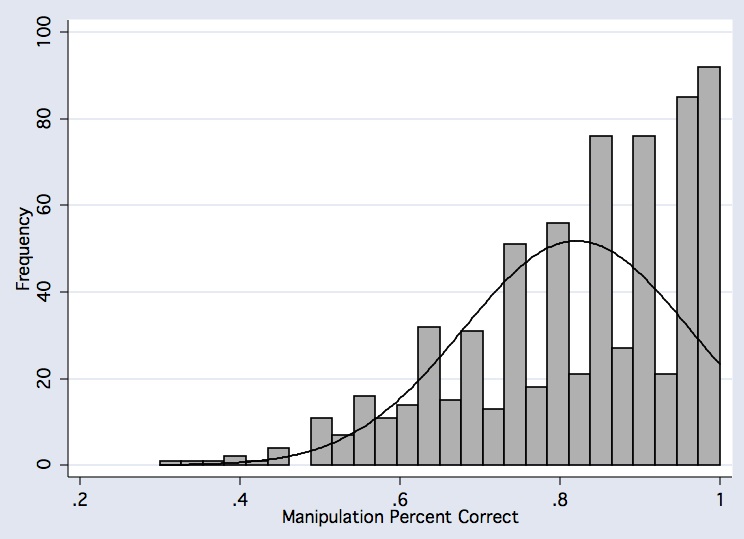
histogram vmnm_manip_mn if comp_useable_vmnm==1, frequency bfcolor(gs11) blcolor(black) normal normopts( clcolor(black) ) xtitle(Manipulation Percent Correct)
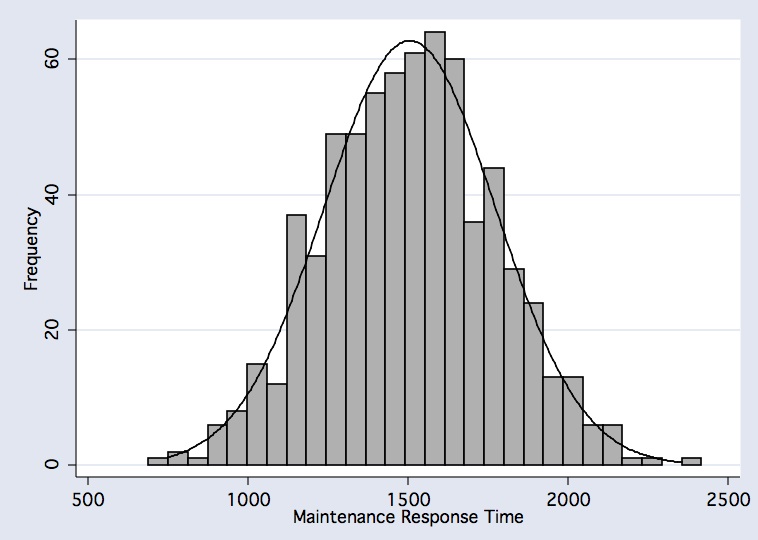
histogram vmnm_main_mnrt if comp_useable_vmnm==1, frequency bfcolor(gs11) blcolor(black) normal normopts( clcolor(black) ) xtitle(Maintenance Response Time)
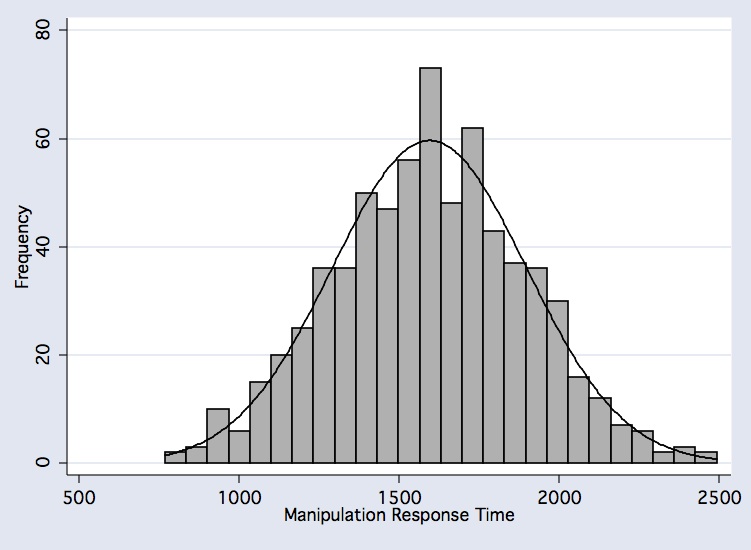
histogram vmnm_manip_mnrt if comp_useable_vmnm==1, frequency bfcolor(gs11) blcolor(black) normal normopts( clcolor(black) ) xtitle(Manipulation Response Time )